For a tour of our active research areas, please click the images below. To see a more general overview of our research interests, please continue to scroll through this page.
Current Funding
 |
 |
 |
Recently Completed Funding
 |
Biophysical chemistry in the Showalter Laboratory
Molecular biophysics, or biophysical chemistry depending on your point of view, is a merging of the biological, chemical, and physical scientific cultures. Our overall goal is to provide fundamental insight into the systems and processes from biochemistry, using the principles and quantitative laws from physical chemistry. Towards this goal, we pursue the following general aims:
- To establish NMR methodology enabling quantitative and comprehensive structural biology for molecules that are traditionally under-represented in structural databanks.
- Provide predictive connections between structure and function for highly flexible biomolecules, which are now agreed to be very common.
The model systems we employ in our research are generally proteins or nucleic acids that function in regulating eukaryotic gene expression. We are particularly interested in the molecular mechanism of transcription control by intrinsically disordered proteins. Our research efforts span the range from basic science and methods development through research with potential biomedical applications.
For a tour of techniques frequently used in our laboratory, please continue reading below.
Unconventional NMR used routinely in our laboratory
Research made possible by: Lloyd Jackman NMR Facility
We do not take the same approach to biomolecular NMR spectroscopy as the majority of laboratories, because we are not focused on the cooperatively folded protein systems that conventional techniques were designed for.
Please follow this link for a more detailed discussion of our carbon-detected biomolecular NMR strategy. (coming soon)
Rigorous thermodynamic characterization of biomolecular interaction mechanisms
Research made possible by: Huck Automated Biological Calorimetry Facility
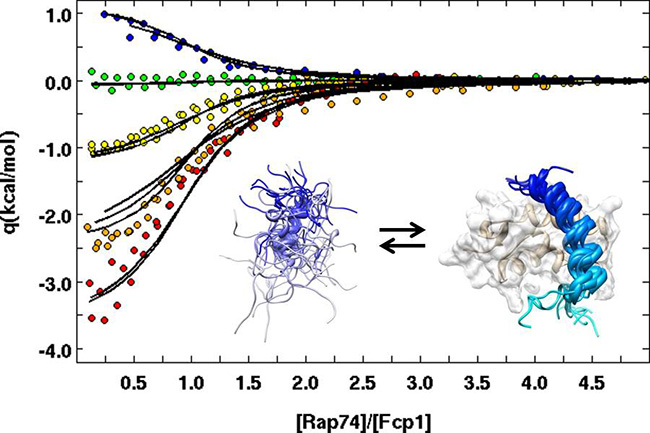
Biomolecular calorimetry is a mainstay in our laboratory, because it offers the greatest potential to thoroughly characterize the thermodynamics of biomolecular interactions and folding transitions, while doing so in as compact a set of experiments as possible.
Please follow this link for a discussion of our temperature-dependent ITC fitting strategy. (coming soon)
An eye for biological application
Research made possible through collaboration with Dr. Melissa Rolls.
We feel it is important for new biophysical knowledge to generate predictive models that lead to better understanding of biological outcomes. Motivated by this core principle, we perform biological assays within our laboratory or in collaboration with our PSU colleagues whenever possible. In addition to our biochemistry laboratory we maintain our own mammalian cell culture room. Just to give one example of the opportunities available at Penn State, through collaboration with colleagues like Dr. Rolls, we are able to perform state-of-the-art cellular imaging assays.