(Undergraduate authors denoted in Penn State blue)
2024
206. McKinley, L. N., Meyer, M. O., Sebastian, A., Chang, B. K., Messina, K. J., Albert, I., Bevilacqua, P. C. Direct testing of natural twister ribozymes from over a thousand organisms reveals a broad tolerance for structural imperfections. bioRxiv (2024). [PubMed]

205. Forstmeier, P. C., Meyer, M. O., Bevilacqua, P. C. The Functional RNA IDentification (FRID) pipeline: Identification of potential pseudoknot-containing RNA elements as therapeutic targets for SARS-CoV-2. bioRxiv (2024). [PubMed]. 
204. Douds, C. A.; Babitzke, P.; Bevilacqua, P. C. A new reagent for in vivo structure probing of RNA G and U residues that improves RNA structure prediction alone and combined with DMS. RNA 30, 901-919 (2024). [PubMed]

203. Jolley, E. A., Bevilacqua, P. C. Single-cell probing of RNA structure. Nat. Methods, 21, 377-378 (2024). [PubMed] (link for full-text)

202. McKinley, L. N., Kern, R. G., Assmann, S. M. & Bevilacqua, P. C. Flanking sequence cotranscriptionally regulates twister ribozyme activity. Biochemistry 63, 53-68 (2024). [PubMed] (link to 50 free e-prints).

2023
201. Sieg, J. P., Jolley, E. A., Huot, M. J., Babitzke, P., Bevilacqua, P. C. In vivo-like nearest neighbor parameters improve prediction of fractional base-pairing in cells. Nucleic Acids Res. 51, 11298-11317 (2023). [PubMed]

200. Meyer M.O., Yamagami R., Choi S., Keating C.D., Bevilacqua P.C. RNA folding studies inside peptide-rich droplets reveal roles of modified nucleosides at the origin of life. Sci. Adv. 22, 9(38):eadh5152 (2023). [PubMed].

199. Williams, A. M., Jolley, E. A., Santiago-Martínez, M. G., Chan, C. X., Gutell, R. R., Ferry, J. G., Bevilacqua, P. C. In vivo structure probing of RNA in Archaea: Novel insights into the ribosome structure of Methanosarcina acetivorans. RNA 29, 1610-1620 (2023). [PubMed]

198. Meyer, M. O., Choi, S., Keating, C. D., Bevilacqua, P. C. & Yamagami, R. Structure-seq of tRNAs and other short RNAs in droplets and in vivo. Methods Enzymol. 681, 81-126 (2023). [PubMed].

197. Jolley, E. A., Yakhnin, H., Tack, D. C., Babitzke, P. & Bevilacqua, P. C. Transcriptome-wide probing reveals RNA thermometers that regulate translation of glycerol permease genes in Bacillus subtilis. RNA 29, 1368-1378 (2023). [PubMed].

196. Sieg J.P., Arteaga S.J., Znosko B.M., Bevilacqua PC. Meltr software provides facile determination of nucleic acid thermodynamics. Biophys Rep 3, 100101 (2023). [PubMed].

195. Assmann S.M., Chou H.L., Bevilacqua P.C. Rock, scissors, paper: How RNA structure informs function. Plant Cell 35, 1671-1707 (2023). [PubMed].

2022
194. Sieg, J. P., McKinley, L. N., Huot, M. J., Yennawar, N. H., Bevilacqua, P. C. The metabolome weakens RNA thermodynamic stability and strengthens RNA chemical stability. Biochemistry 61, 2579-2591 (2022). [PubMed].

193. Bevilacqua, P. C., Tolbert, B. S. Regulatory mechanisms through RNA conformational switching and dynamics, J. Mol. Biol. 434, 167794 (2022). [PubMed].

192. Jolley, E. A., Bormes, K. M., Bevilacqua, P. C. Upstream flanking sequence assists folding of an RNA thermometer. J. Mol. Biol. 434, 167786 (2022). [PubMed].

191. Williams, A. M., Dickson, T. M., Lagoa-Miguel, C. A., Bevilacqua, P. C. Biological solution conditions and flanking sequence modulate LLPS of RNA G-quadruplex structures. RNA 28, 1197-1209 (2022). [PubMed].

190. Yamagami, R., Sieg, J., Assmann, S. A., Bevilacqua, P. C. Genome-wide analysis of the in vivo tRNA structurome reveals RNA structural and modification dynamics under heat stress. Proc. Natl. Acad. Sci. 119, e2201237119 (2022). [PubMed].

189. Ferrero-Serrano, Á., Sylvia, M. M., Forstmeier, P. C., Olson, A. J., Ware, D., Bevilacqua, P. C., Assmann, S. M. Experimental demonstration and pan-structurome prediction of climate associated riboSNitches in Arabidopsis. Genome Biol. 23, 1-28 (2022). [PubMed].

188. Choi, S., Meyer, M. O., Bevilacqua, P. C., Keating, C. D. Phase-specific RNA accumulation and duplex thermodynamics in multiphase coacervate models for membraneless organelles. Nature Chem. 14, 1110-1117 (2022). [PubMed].
187. Bevilacqua, P. C., Williams, A. M., Chou, H. L., Assmann, S. M. RNA multimerization as an organizing force for liquid-liquid phase separation. RNA 28, 16-26 (2022). [PubMed]. 
2021
186. Poudyal, R. R., Sieg, J. P., Portz, B., Keating, C. D., Bevilacqua, P. C. RNA sequence and structure assembly and function of RNA condensates. RNA 12, 1589-1601 (2021). [PubMed].

185. Veenis, A. J., Li, P., Soudackov, A. V., Hammes-Schiffer, S., Bevilacqua, P. C. Investigation of the pKa of the nucleophilic O2′ of the hairpin ribozyme. J. Phys. Chem. B. 43, 11869-11883 (2021). [PubMed]. CLICK HERE TO SEE DREW’S COVER ART!!

184. Williams, A. M., Poudyal, R., Bevilacqua, P. C. Long tracts of guanines drive aggregation of RNA G-quadruplexes in the presence of spermine. Biochemistry 60, 2715-2726 (2021). [PubMed].

183. Yamagami, R., Sieg, J. P., Bevilacqua, P. C. Functional roles of chelated magnesium ions in RNA folding and function. Biochemistry 60, 2374-2386 (2021). [PubMed]

182. Kayedkhordeh, M., Yamagami, R., Bevilacqua, P. C., Mathews, D. H. Inverse RNA folding workflow to design and test ribozymes including pseudoknots. Methods Mol. Biol. 2167, 113-143 (2021). [PubMed]

181. Poudyal, R. R., Meyer, M. O., Bevilacqua, P. C. Measuring the activity and structure of functional RNAs inside compartments formed by liquid-liquid phase separation. Methods Enzymol. 646, 307-327 (2021). [PubMed]

2020
180. Namani, T., Snyder, S., Eagan, J., Bevilacqua, P. C., Wesdemiotis, C. & Sahai, N. Amino Acid Specific Nonenzymatic Montmorillonite-Promoted RNA Polymerization. ChemSystemsChem (2020). [Link]

179. Cakmak, F. P., Choi, S., Meyer, M. O., Bevilacqua, P. C., Keating, C. D. Prebiotically-relevant low polyion multivalency can improve functionality of membraneless compartments. Nature Commun. 11, 5949 (2020). [PubMed]
178. Renda, A., Poly, S., Lai, Y. J., Pannuri, A., Yakhnin, H., Potts, A. H., Bevilacqua, P. C., Romeo, T. & Babitzke, P. CsrA-Mediated Translational Activation of ymdA Expression in Escherichia coli. mBio 11 (2020). [PubMed].

177. Ritchey, L. E., Tack, D. C., Yakhnin, H., Jolley, E. A., Assmann, S. M., Bevilacqua, P. C., Babitzke, P. Structure-seq2 probing of RNA structure upon amino acid starvation reveals both known and novel RNA switches in Bacillus subtilis. RNA 26, 1431-1447 (2020). [PubMed]

176. Tack, D. C.; Su, Z.; Yu, Y.; Bevilacqua, P. C.; Assmann, S. M. Tissue-specific changes in the RNA structurome mediate salinity response in Arabidopsis. RNA 26, 492-511 (2020). [PubMed].

2019
175. Waldron, J. A., Tack, D. C., Ritchey, L. E., Gillen, S. L., Wilczynska, A., Turro, E., Bevilacqua, P. C., Assmann, S. M., Bushell, M., Le Quesne, J. mRNA structural elements immediately upstream of the start codon dictate dependence upon eIF4A helicase activity. Genome Biol 20, 300 (2019). [PubMed].

174. Messina, K. J., Kierzek, R., Tracey, M. A., Bevilacqua, P. C. Small molecule rescue and glycosidic conformational analysis of the twister ribozyme. Biochemistry 58, 4857-4868 (2019). [PubMed].

173. Leamy, K. A., Yamagami, R., Yennawar, N., Bevilacqua, P. C. Single nucleotide control of tRNA folding cooperativity. Proc Natl. Acad. Sci. 116, 23075-23082 (2019). [PubMed].

172. Yamagami, R., Huang, R. & Bevilacqua, P. C. Cellular concentrations of nucleotide diphosphate-chelated magnesium ions accelerate catalysis by RNA and DNA enzymes. Biochemistry 58, 3971-3979 (2019). [PubMed]

171. Mitchell, D., Assmann, S. M., Bevilacqua, P. C. Probing RNA structure in vivo. Curr. Opin. Struct. Biol. 59, 151-158 (2019). [PubMed].

170. Poudyal, R., Keating, C. D., Bevilacqua, P. C. Polyanion-assisted catalysis inside complex coacervates. ACS Chem. Biol. 14, 1243-1248 (2019). [PubMed]

169. Bevilacqua, P. C., Harris, M. E., Piccirilli, J. A., Gaines, C., Ganguly, A., Kostenbader, K., Ekesan, S., York D. M. An ontology for facilitating discussion of catalytic strategies of RNA-cleaving enzymes. ACS Chem. Biol. 14, 1068-1076 (2019) [PubMed].

168. Ritchey, L.E., Su, Z., Assmann, S.M., Bevilacqua, P.C. In vivo genome-wide RNA structure probing with Structure-seq2. Methods Mol. Biol. 1933, 305-341 (2019) [PubMed].

167. Poudyal, R.R., Guth, R.M., Veenis, A.J., Frankel, E.A., Keating, C.D., Bevilacqua, P.C. Template-directed RNA polymerization and enhanced ribozyme catalysis inside membraneless compartments formed by coacervates. Nat. Commun. 10, 490 (2019) [PubMed] (open access).

166. Yamagami, R., Kayedkhordeh, M., Mathews, D.H., Bevilacqua, P.C. Design of highly active double-pseudoknotted ribozymes: a combined computational and experimental study. Nucleic Acids Res. 47, 29-42 (2019) [PubMed] (open access).

165. Mitchell III, D., Renda, A.J., Douds, C.A, Babitzke, P., Assmann, S.M., Bevilacqua, P.C. In vivo structural probing of uracil and guanine base pairing by 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) RNA 25, 147-157 (2019) [PubMed].
 2018
2018
164. Su, Z., Tang, Y., Ritchey, L.E., Tack, D.C., Zhu, M., Bevilacqua, P. C., Assmann, S. M. Genome-wide RNA structurome reprogramming reveals temperature-dependent regulation. Proc. Natl. Acad. Sci. 115, 12170-12175 (2018) [PubMed] (open access).

163. Bevilacqua, P.C. & Assmann, S.M. Technique development for probing RNA structure in vivo and genome-wide. Book Chapter in RNA World 5th Editors: J. Atkins, T.R. Cech, J. A. Steitz. (2018). [PubMed] (link to 50 free e-prints).

162. Messina, KJ & Bevilacqua, PC. Cellular small molecules contribute to twister ribozyme catalysis. J. Am. Chem. Soc. 140, 10578-10582 (2018) [PubMed] (link to 50 free e-prints).
161. Bevilacqua, PC, Bingaman, JL, Frankel, E A, Messina, K J, Seith, D D Key catalytic strategies in ribozymes, Chapter in “Catalysis in Chemistry and Biology”, Proceedings of the 24th International Solvay Conference in Chemistry: Catalysis in Chemistry and Biology. (2018) [Google Books].

Speakers in Section on Catalysis by Ribozymes in Molecular Machines (L to R): Phil Bevilacqua, Dan Herschlag, David Lilley, John Christodoulou, Ron Breaker, Marina Rodnina, Darrin York.
160. Yamagami R, Bingaman JL, Frankel EA, Bevilacqua PC. Cellular conditions of weakly chelated magnesium ions strongly promote RNA stability and catalysis. Nat. Commun. 9, 2149 (2018) [PubMed] (open access).

159. Leamy KA, Yennawar NH, Bevilacqua PC. Molecular mechanism for folding cooperativity of functional RNAs in living organisms. Biochemistry 57, 2994-3002 (2018). [PubMed] (link to 50 free e-prints).

158. Poudyal RR, Pir Cakmak F, Keating CD, Bevilacqua PC. Physical principles and extant biology reveal roles for RNA-containing membraneless compartments in origins of life chemistry. Biochemistry 57, 2509-2519 (2018) [PubMed] (link to 50 free e-prints).
157. Bevilacqua, P. C., Assmann, S. M. RNA structure: A LASER-focused view into cells, Nat. Chem. Biol. 14, 200-201 (2018) [PubMed].

156. Tack, D. C., Tang, Y., Ritchey, L. E., Assmann, S. M., Bevilacqua, P. C. “StructureFold2: Bringing chemical probing data into the computational fold of RNA structural analysis.” Methods 143, 12-15 (2018) [PubMed].

155. Frankel, E.A. & Bevilacqua, P.C. “Complexity in pH-dependent ribozyme kinetics: Dark pKa shifts and wavy rate-pH profiles.” Biochemistry 57, 483-488 (2018) [PubMed] (link to 50 free e-prints)

154. Seith, D., Bingaman, J. L., Veenis, A.J., Button, A.C., & Bevilacqua, P. C. “Elucidation of catalytic strategies of small nucleolytic ribozymes from comparative analysis of active sites.” ACS Catalysis 8, 314-327 (2018) [link] [Minor corrections and additions].

153. Spasic, A., Assmann, S. M., Bevilacqua, P. C., & Mathews, D. H. “Modeling RNA secondary structure folding ensembles using SHAPE mapping data.” Nucleic Acids Res. 46. 314-323 (2018) [PubMed].

152. Mitchell, D. III, Ritchey, L. E., Park, H., Babitzke, P., Assmann, S. A., & Bevilacqua, P. C. “Glyoxals as in vivo RNA structural probes of guanine base pairing.” RNA 24, 114-124 (2018). [PubMed]
2017
151. Bingaman, J. L., Gonzalez, I. Y., Wang, B. & Bevilacqua, P. C. “Activation of the glmS ribozyme nucleophile via overdetermined hydrogen bonding.” Biochemistry 56, 4313-4317 (2017). [PubMed] (link to 50 free e-prints)
150. Leamy, K. A., Yennawar, N. H. & Bevilacqua, P. C. “Cooperative RNA folding under cellular conditions arises from both tertiary structure stabilization and secondary structure destabilization.” Biochemistry 56, 3422-3433 (2017). [PubMed] (link for 50 free e-prints)

149. Ritchey, L. E., Su, Z., Tang, Y., Tack, D. C., Assmann, S. M. & Bevilacqua, P. C. “Structure-seq2: Sensitive and accurate genome-wide profiling of RNA structure in vivo.” Nucleic Acids Res. 45, e135 (2017). [PubMed]

148. Frankel, E. A., Strulson, C., Keating, C. D. & Bevilacqua, P. C. “Cooperative interactions in the hammerhead ribozyme drive pKa shifting of G12 and its stacked base C17.” Biochemistry 56, 2537-2548 (2017). [PubMed]
(link for 50 free e-prints)
147. Bingaman, J. L., Messina, K. J., Bevilacqua, P. C. “Probing fast ribozyme reactions under biological conditions with rapid quench-flow kinetics.” Methods 120, 125-134 (2017). [PubMed]

146. Bingaman, J. L., Zhang, S., Stevens, D. R., Yennawar, N. H., Hammes-Schiffer, S., Bevilacqua, P. C. “GlcN6P Cofactor serves multiple catalytic roles in the glmS ribozyme.” Nat. Chem. Biol. 13, 439-445 (2017). [PubMed]

2016
145. Bingaman, J. L., Frankel, E. A., Hull, C. M., Leamy, K. A., Messina, K. J., Mitchell, D., 3rd, Park, H., Ritchey, L.E., Babitzke, P., Bevilacqua, P. C. “Eliminating blurry bands in gels with a simple cost-effective repair to the gel cassette.” RNA 22, 1929-1930 (2016). [PubMed]

144. Zhang, S., Stevens, D. R., Goyal, P., Bingaman, J., Bevilacqua, P.C. , Hammes-Schiffer, S. “Assessing the potential effects of active site Mg2+ ions in the glmS ribozyme-cofactor complex” J. Phys. Chem. Letters 7 , 3984-3988 (2016). [PubMed]

143. Bevilacqua, P. C., Ritchey, L. E., Su, Z., Assmann, S. M. “Genome-wide analysis of RNA secondary structure.” Annu. Rev. Genet. 50, 235-266 (2016). [PubMed]

142. Leamy, K. A., Assmann, S. M., Mathews, D. H., Bevilacqua, P. C.. “Bridging the gap between in vitro and in vivo RNA folding.” Q. Rev. Biophys. 49, e10 (2016). [PubMed]

141. Ucisik, M. N., Bevilacqua, P. C., Hammes-Schiffer, S. “Molecular dynamics study of env22 twister ribozyme: Role of Mg2+ ions and hydrogen-bonding network in active site”. Biochemistry 55, 3834-3846. [PubMed]

140. Hull, C. M., Bevilacqua, P. C. “Discriminating self from non-self by RNA: Roles for RNA structure, misfolding, and modification in regulating the innate immune sensor PKR”. Acc. Chem. Res. 49, 1242-1249 (2016). [PubMed]

139. Sherlock, M. E., Rumble, C. A., Kwok, C. K., Breffke, J., Maroncelli, M., Bevilacqua, P. C. “Steady-state and time-resolved studies into the origin of the intrinsic fluorescence of G-quadruplexes.” J. Phys. Chem. B 120, 5146-5158 (2016). [PubMed]

138. Hull, C. M., Anmangandla, A., Bevilacqua, P. C. “Bacterial riboswitches and ribozymes potently activate the human innate immune sensor PKR” ACS Chem. Biol. 11, 1118-1127 (2016). [PubMed]
HIGHLIGHT featured in F1000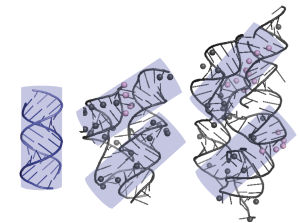
137. Frankel, E. A., Bevilacqua, P. C., Keating, C. D. “Polyamine/nucleotide coacervates provide strong compartmentalization of Mg2+, nucleotides, and RNA” Langmuir 32, 2041-2049 (2016). [PubMed]
136. Yennawar, N. H., Fecko, J. A., Showalter, S. A., Bevilacqua, P. C. “A high-throughput biological calorimetry core—Steps to startup, run, and maintain a multi-user facility” Methods Enzymol. 567, 435-460 (2016). [PubMed]
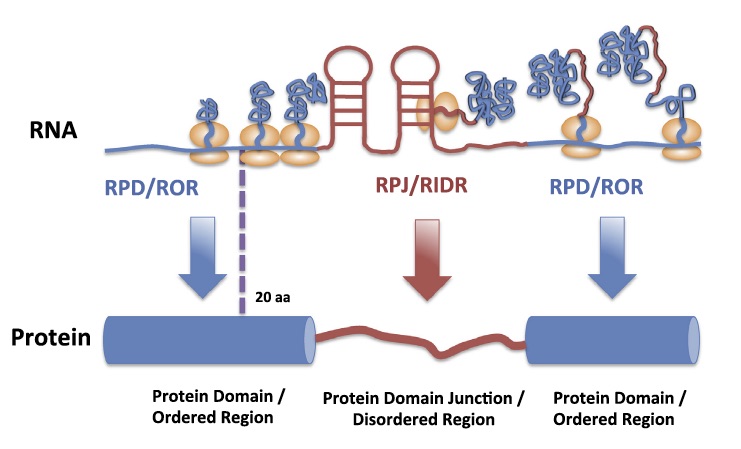
135. Tang, Y., Assmann, S. M., Bevilacqua, P. C. “Protein structure is related to RNA structural reactivity in vivo” Accepted November 2015. Awaiting publication in 2016 in special issue “Challenges in RNA Structural Modeling and Design”. J. Mol. Biol. 428, 758-766 (2016). [PubMed]
HIGHLIGHT featured in F1000
Part of JMB Challenges in RNA Structural Modeling and Design Issue
2015
134. Crenshaw, E., Leung, B. P., Kwok, C. K., Olson, K., Sebastian, N. P., Ansaloni, S., Schweitzer-Stenner, R., Akins, M. R., Bevilacqua, P. C., Saunders, A. J. “Amyloid Precursor Protein Translation is Regulated by a 3’UTR Guanine Quadruplex” PLoS One 10, e0143160, 1-18 (2015). [PubMed]

133. Hull, C. M., Bevilacqua, P. C. “Mechanistic analysis of activation of the innate immune sensor PKR by bacterial RNA” J. Mol. Biol. 427, 3501-3515 (2015). [PubMed]

132. Tang, Y., Bouvier, E., Ding, Y., Nekrutenko, A., Bevilacqua, P.C. Assmann, S. M. “StructureFold: Genome-wide RNA secondary structure mapping and reconstruction in vivo” Bioinformatics 31, 2668-2675. [PubMed]
StructureFold is freely available as a component of Galaxy available at: https://usegalaxy.org/.

131. Ding. Y., Kwok, C. K., Tang, Y., Bevilacqua, P. C., Assmann, S. M. “Structure-Seq: Genome-wide profiling of in vivo RNA structure at single nucleotide resolution” Nat. Protoc. 10, 1050-1066 (2015). [PubMed]

130. Thaplyal, P., Ganguly, A., Hammes-Schiffer, S., Bevilacqua, P. C. “Inverse thio effects in the hepatitis delta virus (HDV) ribozyme reveal that the reaction pathway is controlled by metal ion charge density” Biochemistry 54, 2160–2175 (2015). [PubMed]

129. Kwok, C.K., Tang, Y., Assmann, S. M., Bevilacqua, P. C. “The RNA structurome: transcriptome-wide structure probing with next-gen sequencing” Trends Biochem. Sci. 40, 221-232 (2015). [PubMed]
HIGHLIGHT featured on TiBS cover
128. Bevilacqua, P. C. “The wonder of RNA: A personal reflection of the last 20 Years ”. RNA 21, 515-516 (2015). [PubMed] (pdf)

127. Kwok, C. K., Ding, Y., Shahid, S. Assmann, S. M., and Bevilacqua, P. C. “A stable RNA G-quadruplex within the 5’UTR of Arabidopsis thaliana ATR mRNA downregulates translation” Biochem. J. 467, 91-102 (2015). [PubMed]

126. Zhang, S., Ganguly, A., Goyal, P., Bingaman, J. L., Bevilacqua, P. C., Hammes-Schiffer, S. “Role of the active site guanine in the glmS ribozyme self-cleavage mechanism: quantum mechanical/molecular mechanical free energy simulations” J. Am. Chem. Soc. 137, 784-798 (2015). [PubMed]
HIGHLIGHT featured in JACS Spotlight

2014
125. Thaplyal, P. and Bevilacqua, P.C. “Experimental approaches for measuring pKa’s in RNA and DNA” Methods Enzmol. 549, 188-219 (2014). [PubMed]
124. Dewey, D. C. Strulson, C. A., Cacace, D. N., Bevilacqua, P. C., and Keating, C. D. “Bioreactor droplets from liposome-stabilized all-aqueous emulsions.” Nature Comm. 5, 4670 (2014). [PubMed]

123. Ganguly, A., Thaplyal, P., Rosta, E., Bevilacqua, P. C., and Hammes-Schiffer, S. “Quantum mechanical/molecular mechanical free energy simulations of the self-cleavage reaction in the hepatitis delta virus ribozyme.” J. Am. Chem. Soc. 136, 1483-1496 (2014). [PubMed]
HIGHLIGHT featured in JACS Spotlight

122. Ding, Y., Tang, Y., Kwok, C.K., Zhang, Y., Bevilacqua, P.C. & Assmann, S.M. “In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features.” Nature 505, 696-700 (2014). [PubMed]
HIGHLIGHT articles in Nature, Nature Methods, Nature Chemical Biology, and F1000.

121. Strulson, C. A., Boyer, J. A., Whitman, E. E. & Bevilacqua, P. C. “Molecular crowders and cosolutes promote folding cooperativity of RNA under physiological ionic conditions.” RNA 20, 331-347 (2014). [PubMed]

2013
120. Kwok, C. K., Ding, Y., Tang, Y., Assmann, S. M. & Bevilacqua, P. C. “Determination of in vivo RNA structure in low-abundance transcripts.” Nature Commun. 4, 2971 (2013). [PubMed]

119. Strulson, C.A., Yennawar, N.H., Rambo, R.P., and Bevilacqua, P.C. “Molecular crowding favors reactivity of a human ribozyme under physiological ionic conditions.” Biochemistry, 52, 8187-8197 (2013). [Pubmed]
HIGHLIGHT featured on Biochemistry site for 1 month

118.Wilcox, J.L. and Bevilacqua, P.C. “pKa Shifting in double-stranded RNA (dsRNA) is highly dependent upon nearest neighbors and bulge positioning.” Biochemistry, 52, 7470-7476 (2013). [PubMed]

117.Thaplyal, P., Ganguly, A., Golden, B.L., Hammes-Schiffer, S. and Bevilacqua, P.C. “Thio effects and an unconventional metal ion rescue in the genomic hepatitis delta virus ribozyme.” Biochemistry 52, 6499-6514 (2013). [PubMed]

116. Kwok, C. K., Sherlock, M. E., and Bevilacqua, P. C. “Effect of Loop Sequence and Loop Length on the Intrinsic Fluorescence of G-Quadruplexes.” Biochemistry 52, 3019-3021 (2013). [PubMed]

115. Wilcox, J.L and Bevilacqua, P.C. “A simple fluorescence method for pKa determination in RNA and DNA reveals highly shifted pKa’s” J. Am. Chem. Soc. 135, 7390-7393 (2013). [PubMed]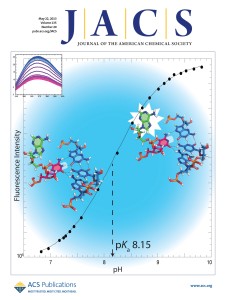
HIGHLIGHT featured in JACS Spotlight, JACS cover.

114. Golden B.L., Hammes-Schiffer S., Carey P.R., Bevilacqua PC. “An integrated picture of HDV ribozyme catalysis.” Springer 2013. Rick Russell editor. pp 135-168. [Springer]

113. Nallagatla, S.R., Jones, C.N., Ghosh, S.K.B., Sharma, S.D., Cameron, C.E., Spremulli, L.L., and Bevilacqua, P.C. “Native tertiary structure and nucleoside modifications suppress tRNA’s intrinsic ability to activate the innate immune sensor PKR.” PLOS ONE 8, e57905, 1-10 (2013). [PubMed]

112. Kwok, C.K., Ding, Y., Sherlock, M.E., Assmann, S.M., Bevilacqua, P.C. “A hybridization-based approach for quantitative and low-bias single-stranded DNA ligation.” Anal Biochem 435, 181-186 (2013). [PubMed].

HIGHLIGHT featured on cover of Anal Biochem
111. Tubbs, J.D., Condon, D.E., Kennedy, S.D., Hauser, M., Bevilacqua, P.C., Turner, D.H. “The nuclear magnetic resonance of CCCC RNA reveals a right-handed helix and revised parameters for AMBER force field torsions improve structural predictions from molecular dynamics.” Biochemistry 52, 996-1010 (2013). [PubMed].
HIGHLIGHT featured on Biochemistry site for 1 month

110. Chen, J., Ganguly, A., Miswan, Z., Hammes-Schiffer, S., Bevilacqua, P.C., Golden, B.L. “Identification of the catalytic Mg2+ ion in the hepatitis delta virus ribozyme”Biochemistry 52, 557-567 (2013). [Pubmed]
HIGHLIGHT featured on Biochemistry site for 1 month

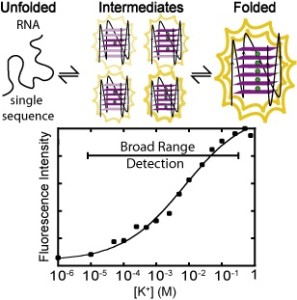
109. Kwok, C.K., Sherlock, M.E., Bevilacqua, P.C. “Decrease in RNA folding cooperativity by deliberate population of intermediates in RNA G-quadruplexes” Angew. Chem. Int. Ed. Engl. 52, 683-686 (2013). [PubMed]
HIGHLIGHT in F1000
2012
108. Patel, S., Blose, J. M., Sokoloski, J. E., Pollack, L., Bevilacqua, P. C. “Specificity of the double-stranded RNA-binding domain from the RNA-activated protein kinase PKR for double-stranded RNA: insights from thermodynamics and small-angle X-ray scattering” Biochemistry 51, 9312-9322 (2012). [Pubmed]
107. Heinicke, L. A., Bevilacqua, P. C. “Activation of PKR by RNA misfolding: HDV ribozyme dimers activate PKR.” RNA 18, 2157-2165 (2012). [Pubmed]
106. Bevilacqua, P. C., Breen, P. C., Thaplyal, P. “Prospecting for aptamers in the human genome” Chemistry & Biology, 19, 1218-1220 (2012). [Pubmed]
105. Strulson, C. A., Molden, R. C., Keating, C. D., Bevilacqua, P. C. “RNA catalysis through compartmentalization” Nature Chemistry 4, 941-946 (2012).[Pubmed]
HIGHLIGHT in phys.org, Science Daily, and F1000
104. Toroney, R., Hull, C. M., Sokoloski, J. E., Bevilacqua, P. C. “Mechanistic characterization of the 5′-triphosphate-dependent activation of PKR: Lack of 5′-end nucleobase specificity, evidence for a distinct triphosphate binding site, and a critical role for the dsRBD” RNA 18, 1862-1874 (2012). [Pubmed]
103. Sokoloski, J. E., Bevilacqua, P. C. “Analysis of RNA folding and ligand binding by conventional and high-throughput calorimetry.” Methods Mol. Biol. 905, 145-74 (2012). [Pubmed]
102. Chadalavada, D. M., Cerrone-Szakal, A. L., Wilcox, J. L., Siegfried, N. A., Bevilacqua, P. C. “Mechanistic analysis of the Hepatitis Delta Virus (HDV) ribozyme: Methods for RNA preparation, structure mapping, solvent isotope effects, and co-transcriptional cleavage.” Methods Mol. Biol. 848, 21-40 (2012). [Pubmed]
101. Mullen, M. A., Assmann, S. M., Bevilacqua, P. C. “Toward a digital gene response: RNA G-quadruplexes with fewer quartets fold with higher cooperativity.” J. Am. Chem. Soc. 134, 812-815 (2012). [Pubmed]
100. Sokoloski, J. E., Dombrowski, S. E., Bevilacqua, P. C. “Thermodynamics of ligand binding to a heterogeneous RNA population in the malachite green aptamer.” Biochemistry 51, 565-572 (2012). [Pubmed]
2011
99. Wilcox, J. L., Ahluwalia, A. K., Bevilacqua, P. C. “Charged nucleobases and their potential for RNA catalysis.” Acc. Chem. Res. 44, 1270-1279 (2011). [Pubmed]
98. Ganguly, A., Bevilacqua, P. C., Hammes-Schiffer, S. “Quantum Mechanical/Molecular Mechanical study of the HDV ribozyme: Impact of the catalytic metal ion on the mechanism.” J. Phys. Chem. Lett. 2, 2906-2911 (2011). [Pubmed]
97. Sokoloski, J. E., Godfrey, S. A., Dombrowski, S. E., Bevilacqua, P. C. “Prevalence of syn nucleobases in the active sites of functional RNAs.” RNA 17, 1775-1787 (2011). [Pubmed]
HIGHLIGHT in F1000
96. Veeraraghavan, N., Ganguly, A., Golden, B. L., Bevilacqua, P. C., and Hammes-Schiffer, S. “Mechanistic strategies in the HDV ribozyme: Chelated and Diffuse Metal ion Interactions and Active Site Protonation.” J. Phys. Chem. B. 115, 8346-8357 (2011). [Pubmed]
95. Nallagatla, S. R., Toroney, R., Bevilacqua, P. C. “Regulation of innate immunity through RNA structure and the protein kinase PKR” Curr. Opin. Struct. Biol. 21, 119-127 (2011). [Pubmed]
94. Veeraraghavan, N., Ganguly, A., Chen, J. H., Bevilacqua, P. C., Hammes-Schiffer, S., and Golden, B. L. “A metal binding motif in the active site of the HDV ribozyme binds divalent and monovalent ions.” Biochemistry 50, 2672-2682 (2011). [Pubmed]
93. Heinicke, L., Nallagatla, S. R., Hull, C. M., Bevilacqua, P. C. “RNA helical imperfections regulate activation of the protein kinase PKR: Effects of bulge position, size, and geometry.” RNA 17, 957-966 (2011). [Pubmed]
2010
92. Mullen, M. A., Olson, K. J., Dallaire, P., Major, F., Assmann, S. M., Bevilacqua, P. C. “RNA G-quadruplexes in the model plant species Arabidopsis thaliana: Prevalence and possible functional roles.” Nucleic Acids Res. 38 8149-8163 (2010). [Pubmed]
91. Veeraraghavan, N., Bevilacqua, P. C., Hammes-Schiffer, S. “Long distance communication in the HDV ribozyme: Insights from molecular dynamics and experiments.” J. Mol. Biol. 402, 278-291 (2010). [Pubmed]
90. Chen, J.-H., Yajima, R., Chadalavada, D. M., Chase, E., Bevilacqua, P. C., Golden, B. L. “A 1.9 Å crystal structure of the HDV ribozyme pre-cleavage suggests both Lewis acid and general acid mechanisms contribute to phosphodiester cleavage.” Biochemistry 49, 6508-6518 (2010). [Pubmed]
89. Chadalavada, D. M., Gratton, E., and Bevilacqua, P. C. “The human HDV-like CPEB3 ribozyme is intrinsically fast reacting” Biochemistry 49, 5321-5330 (2010). [Pubmed]
88. Toroney, R., Nallagatla, S. R., Boyer, J. A., Cameron, C. E., and Bevilacqua, P. C. “Regulation of PKR by HCV IRES RNA: Importance of Domain II and NS5A.” J. Mol. Biol. 400, 393-412 (2010). [Pubmed]
87. Anderson, B., Muramatsu, H., Nallagatla, S. R., and Bevilacqua, P. C., Weissman, D., Kariko, K. “Incorporation of pseudouridine into mRNA enhances translation by diminishing PKR activation.” Nucleic Acids Res. 38, 5884-5892 (2010). [Pubmed]
86. Siegfried, N. A., Kierzek, R., and Bevilacqua, P. C. “Role of unsatisfied hydrogen bond acceptors in RNA energetics and specificity.” J. Am. Chem. Soc. 132, 5342-5344 (2010). [Pubmed]
85. Siegfried, N. A., O’Hare, B., Bevilacqua, P. C. “Driving forces for nucleic acid pKa shifting: Effects of helix position, temperature, and ionic strength.” Biochemistry 49, 3225-3236 (2010). [Pubmed]
2009
84. Chadalavada, D. M., and Bevilacqua, P. C. “Analyzing RNA and DNA folding using temperature gradient gel electrophoresis (TGGE) with applications to in vitro selections.” Methods Enzymol. 468, 389-408 (2009). [Pubmed]
83. Gong, B., Chen, J.-H., Bevilacqua, P. C., Golden, B. L., and Carey, P. R. “Competition between Co(NH3)63+ and inner sphere Mg2+ ions in the HDV ribozyme.” Biochemistry 48, 11961-11970 (2009). [Pubmed]
82. Toroney, R., and Bevilacqua, P. C. “PKR and the ribosome compete for mRNA.” Nat. Chem. Biol. 5, 873-874 (2009). [Pubmed]
81. Gong, B., Chen, J.-H., Yajima, R., Chen, Y., Chase, E., Chadalavada, D. M., Golden, B. L., Carey, P. R., and Bevilacqua, P. C. “Raman crystallography of RNA.” Methods 49, 101-111 (2009). [Pubmed]
80. Blose, J. M., Lloyd, K. P., and Bevilacqua, P. C. “Portability of the GN(R)A hairpin loop motif between RNA and DNA.” Biochemistry 48, 8787-8794 (2009). [Pubmed]
79. Heinicke, L. A., Toroney, R., and Bevilacqua, P. C. “Evolution of PKR combats viral mimicry.” Cell Sci. 5, 66-74 (2009). [pdf]
78. Heinicke, L. A., Wong, C. J., Lary, J., Nallagatla, S. R., Diegelman-Parente, A., Zheng, X., Cole, J. L., and Bevilacqua, P. C. “RNA dimerization promotes PKR dimerization and activation.” J. Mol. Biol. 390, 319-338 (2009). [Pubmed]
77. Blose, J. M., Proctor, D. J., Misra, V. K., and Bevilacqua, P. C. “Contribution of the closing base pair to exceptional stability in RNA tetraloops: Roles for molecular mimicry and electrostatic factors.” J. Am. Chem. Soc. 131, 8474-8484 (2009). [Pubmed]
76. Siegfried, N. A., & Bevilacqua, P. C. “Thinking inside the box: Designing, implementing, and interpreting thermodynamic cycles to dissect cooperativity in RNA and DNA folding.” Methods Enzymol. 455, 365-393 (2009). [Pubmed]
75. Chen, J.-H., Gong, B., Bevilacqua, P. C., Carey, P. R., and Golden, B. “A catalytic metal ion interacts with the cleavage site GU wobble in the HDV ribozyme.” Biochemistry 48, 1498-1507 (2009). [Pubmed]
HIGHLIGHT in F1000
74. McGraw, A., Mokdad, A., Major, F., Bevilacqua, P. C., and Babitzke, P. “Molecular basis of TRAP-5’SL RNA interaction in the Bacillus subtilis trp operon transcription attenuation mechanism.” RNA 15, 55-66 (2009). [Pubmed]
2008
73. Bevilacqua, P. C., & Russell, R. “Editorial overview: Exploring the vast dynamics of RNA dynamics.” Curr. Opin. Chem. Biol. 12, 601-603 (2008). [Pubmed]
72. Nallagatla, S. R., Toroney, R., Bevilacqua, P.C. “A brilliant disguise for self RNA: 5′-end and internal modifications of primary transcripts suppress elements of innate immunity.” RNA Biology 5, 25-29 (2008). [Pubmed]
71. Cerrone-Szakal, A.L., Siegfried, N.A., Bevilacqua, P.C. “Mechanistic characterization of the HDV genomic ribozyme: Solvent isotope effects and proton inventories in the absence of divalent metal ions support C75 as the general acid.” J. Am. Chem. Soc. 130, 14504-14520 (2008). [Pubmed]
70. Cerrone-Szakal, A.L., Chadalavada, D.M., Golden, B.L., Bevilacqua, P.C. “Mechanistic characterization of the HDV genomic ribozyme: The cleavage site base pair plays a structural role in facilitating catalysis.” RNA 14, 1746-1760 (2008). [Pubmed]
69. Gong, B., Chen, Y., Christian, E.L., Chen, J.H., Chase, E., Chadalavada, D.M., Yajima, R., Golden, B.L., Bevilacqua, P.C., Carey, P.R. “Detection of innersphere interactions between magnesium hydrate and the phosphate backbone of the HDV ribozyme using Raman crystallography.” J. Am. Chem. Soc. 130, 9670-9672 (2008). [Pubmed]
68. Nallagatla, S. R., Bevilacqua, P.C. “Nucleoside modifications modulate activation of the protein kinase PKR in an RNA structure-specific manner” RNA 14, 1201-1213 (2008). [Pubmed]
67. Bevilacqua, P.C., Blose, J.M. “Structures, kinetics, thermodynamics, and biological functions of RNA hairpins.” Annu. Rev. Phys. Chem. 59, 79-103 (2008). [Pubmed]
66. Bevilacqua, P.C. “Proton transfer in ribozyme catalysis.” Ribozymes and RNA Catalysis. D.M. Lilley and F. Eckstein eds. RSC Publishing, Cambridge, UK (2008). [pdf]
2007
65. Nallagatla, S.R., Hwang, J., Toroney, R., Zheng, X., Cameron, C.E., Bevilacqua. P.C. “5′-triphosphate-dependent activation of PKR by RNAs with short stem-loops.” Science 318, 1455-1458 (2007). [Pubmed]
HIGHLIGHT in F1000
64. Gong, B, Chen, J.H., Chase, E., Chadalavada, D.M., Yajima, R., Golden, B.L., Bevilacqua, P.C., Carey, P.R. “Direct measurement of a pKa near neutrality for the catalytic cytosine in the genomic HDV ribozyme by Raman crystallography.” J. Am. Chem. Soc. 129, 13335-13342 (2007). [Pubmed]
63. McGraw, A.P., Bevilacqua, P.C., Babitzke, P. “TRAP-5′ Stem-Loop interaction increases the effect the efficiency of transcription termination in the Bacillus subtillis trpEDCFBA operon leader region by increasing the rate of TRAP binding to the nascent transcript.” RNA 13, 2020-2033 (2007). [Pubmed]
62. Chadalavada, D.M., Cerrone-Szakal, A.L., Bevilacqua, P.C. “Wild-type is the optimal sequence of the HDV ribozyme under co-transcriptional conditions.” RNA 13, 2189-2201 (2007). [Pubmed]
61. Bevilacqua, P.C., SantaLucia, J. Jr. “The biophysics of RNA.” ACS Chem. Biol. 2, 440-444 (2007). [Pubmed]
60. Bevilacqua, P.C., Cerrone-Szakal, AL, Siegfried, N.A. “Insight into the functional diversity of RNA through model-making with applications to data fitting.” Q. Rev. Biophys. 40, 55-85 (2007). [Pubmed]
59. Blose, J.M., Silverman, SK, Bevilacqua, P.C. “A simple molecular model for thermophilic adaptation of functional nucleic acids.” Biochemistry 46, 4232-4240 (2007). [Pubmed]
58. Nakano, S., Bevilacqua, P.C. “Mechanistic characterization of the HDV genomic ribozyme: A mutant of the C41 motif provides insight into the positioning and thermodynamic linkage of metal ions and protons.” Biochemistry 46, 3001-3012 (2007). [Pubmed]
57. Yajima, R., Proctor, D.J., Kierzek, R., Kierzek, E., Bevilacqua, P.C. “A conformationally restricted guanosine analog reveals the catalytic relevance of three structures of an RNA enzyme.” Chem. Biol. 14, 23-30 (2007). [Pubmed]
56. Siegfried, N.A., Metzger, S.L., Bevilacqua, P.C. “Folding cooperativity in RNA and DNA is dependent on position in the helix.” Biochemistry 46, 172-181 (2007). [Pubmed]
2006
55. Bevilacqua, P.C., Yajima, R. “Nucleobase catalysis in ribozyme mechanism.” Curr. Opin. Chem. Biol. 10, 455-464 (2006). [Pubmed]
54. Ma, H., Proctor, D. J., Kierzek, E., Kierzek, R., Bevilacqua, P. C. Gruebele, M. “Exploring the energy landscape of a small RNA hairpin.” J. Am. Chem. Soc. 128, 1523-1530 (2006). [Pubmed]
2005
53. Bevilacqua, P. C., Brown, T. S., Chadalavada, D., Lecomte, J., Moody, E., and Nakano, S.-i. “Linkage between proton binding and folding in RNA: Implications for RNA catalysis.” Biochem. Soc. Trans. 33, 466-470 (2005). [Pubmed]
52. Brown, T. S., and Bevilacqua, P. C. “Method for assigning double-stranded RNA structures.” Biotechniques 38, 368-372 (2005). An overview of the article appears in the front of this issue. [Pubmed]
HIGHLIGHT in Biotechniques
51. Moody, E. M., Lecomte, J. T. J., and Bevilacqua, P. C. “Linkage between proton binding and RNA folding: A thermodynamic framework and its experimental application for investigating pKa shifting.” RNA 11, 157-172 (2005). [Pubmed]
2004
50. Zheng, X., and Bevilacqua, P. C. “Activation of the protein kinase PKR by short double-stranded RNAs with single-stranded tails.” RNA 10, 1934-1945 (2004). [Pubmed]
49. Tian, B., Bevilacqua, P. C., Diegelman-Parente, A., and Mathews, M. B. “The double-stranded RNA binding motif: interference and much more.” Nat. Rev. Mol. Cell. Biol. 5, 1013-1023 (2004). [Pubmed]
48. Proctor, D. J., Ma, H., Kierzek, E., Kierzek, R., Gruebele, M., and Bevilacqua, P. C. “Folding thermodynamics and kinetics of YNMG RNA hairpins: Specific incorporation of 8-bromoguanosine leads to stabilization by enhancement of the folding rate.” Biochemistry 43, 14004-14014 (2004). [Pubmed]
47. Paxon, T. L., Brown, T. S. Hsiao-yu, N. L. Brancato, S. J., Roddy, E. S., Bevilacqua, P. C., and Ewing, A. G. “Continuous monitoring of enzyme reactions on a microchip: Application to catalytic RNA self-cleavage.” Anal. Chem. 76, 6921-6927 (2004). (A 1 page overview of the article appears in the front of this issue.) [Pubmed]
HIGHLIGHT in Analytical Chemistry
46. Moody, E. M., Brown, T. S., and Bevilacqua, P. C. “Simple method for determining nucleobase pKa values by indirect labeling and demonstration of a pKa of neutrality in dsDNA.” J. Am. Chem. Soc. 126, 10200-10201 (2004). [Pubmed]
45. Brown, T. S., Chadalavada, D. M., and Bevilacqua, P. C. “Design of a highly reactive HDV ribozyme sequence uncovers facilitation of RNA folding by alternative pairings and physiological ionic strength.” J. Mol. Biol. 341, 695-712 (2004). [Pubmed]
44. Moody, E. M., and Bevilacqua, P. C. “Structural and energetic consequences of expanding a highly cooperative stable DNA hairpin loop.” J. Am. Chem. Soc. 126, 9570-9577 (2004). [Pubmed]
43. Moody, E. M., Feerrar, J. C., and Bevilacqua, P. C. “Evidence that folding of RNA tetraloop hairpin is less cooperative than its DNA counterpart.” Biochemistry, 43, 7992-7998 (2004). [Pubmed]
42. Bevilacqua, P. C. “Mechanism of catalytic RNA.” Biopolymers 73, 69-70 (2004). [Pubmed]
41. Bevilacqua, P. C., Brown, T. S., Nakano, S., and Yajima, R. “Catalytic roles for proton transfer and protonation in ribozymes.” Biopolymers 73, 90-109 (2004). [Pubmed]
2003
40. Moody, E. M., and Bevilacqua, P. C. “Folding of a stable DNA motif involves a highly cooperative network of interactions.” J. Am. Chem. Soc.. 125, 16285-16293 (2003). [Pubmed]
39. Schaak, J. E., Babitzke, P., and Bevilacqua, P. C. “Phylogenetic conservation of RNA secondary and tertiary structure in the trpEDCFBA operon leader transcript in Bacillus.” RNA 9, 1502-1515 (2003). [Pubmed]
38. Schaak, J. E., Yakhnin, H., Bevilacqua, P. C., and Babitzke, P. “A Mg2+-dependent RNA tertiary structure forms in the Bacillus subtilis trp operon leader transcript and appears to interfere with trpE translation control by inhibiting TRAP binding.” J. Mol. Biol., 332, 555-574 (2003). [Pubmed]
37. Babitzke, P., Schaak, J., Yakhnin, A. V., and Bevilacqua, P. C. “Role of RNA structure in transcription attenuation in Bacillus subtilis: the trpEDCFBA operon as a model system.” Methods Enzymol., 371, 392-404 (2003). [Pubmed]
36. Bevilacqua, P. C., Brown, T. S., Chadalavada, D. M., Parente, A. D. and Yajima, R. “Kinetic analysis of ribozyme cleavage.” In Kinetic Analysis of Macromolecules: A Practical Approach (K. Johnson, ed.) Oxford University Press. Chpt 3, 49-74 (2003). [pdf]
35. Nakano, S., Cerrone, A. L., and Bevilacqua, P. C. “Mechanistic characterization of the HDV genomic ribozyme: Classifying the catalytic and structural metal ion sites within a multichannel reaction mechanism.” Biochemistry, 42, 2982-2994 (2003). [Pubmed]
34. Bevilacqua, P. C. “Mechanistic considerations for general acid-base catalysis by RNA: Revisiting the mechanism of thehairpin ribozyme.” Biochemistry, 42, 2259-2265 (2003). [Pubmed]
33. Proctor, D. J., Kierzek, E., Kierzek, R., and Bevilacqua, P. C. “Restricting the conformational heterogeneity of RNA by specific incorporation of 8-Bromoguanosine.” J. Am. Chem. Soc., 125, 2390-2391 (2003). [Pubmed]
32. Moody, E. M., and Bevilacqua, P. C. “Thermodynamic coupling of the loop and stem in unusually stable DNA hairpins closed by CG base pairs.” J. Am. Chem. Soc., 125, 2032-2033 (2003). [Pubmed]
2002
31. Nakano, M., Moody, E. M., Liang, J., and Bevilacqua, P. C. “Selection for thermodynamically stable DNA tetraloops using temperature gradient gel electrophoresis reveals four motifs: d(cGNNAg), d(cGNABg), d(cCNNGg), and d(gCNNGc).” Biochemistry, 41, 14281-14292 (2002). [Pubmed]
30. Diegelman-Parente, A., and Bevilacqua, P. C. “A mechanistic framework for co-transcriptional folding of the HDV genomic ribozyme in the presence of downstream sequence.” J. Mol. Biol., 324, 1-16 (2002). [Pubmed]
29. Proctor, D. J., Schaak, J. E., Bevilacqua, J. M., Falzone, C. J. and Bevilacqua, P. C. “Isolation and characterization of a family of stable tetraloops with the motif YNMG that participate in tertiary interactions.” Biochemistry, 41, 12062-12075 (2002). [Pubmed]
28. Bevilacqua, P. C. “Battle for the bulge: Directing natural products to DNA defects.” Chem. Biol., 9, 854-855 (2002). [Pubmed]
27. Bevilacqua, P.C. and Turner, D.H. “Use of fluorescence spectroscopy to elucidate RNA folding pathways.” In Current Protocols in Nucleic Acid Chemistry (S. Beaucage et al., eds.). John Wiley & Sons, New York. 11.8.1-11.8.6 (2002). [Pubmed]
26. Chadalavada, D. M., Senchak, S. E., and Bevilacqua, P. C. “The folding pathway of the genomic hepatitis delta virus ribozyme is dominated by slow folding of the pseudoknots.” J. Mol. Biol., 317, 559-575 (2002). [Pubmed]
2001
25. Nakano, S. and Bevilacqua, P. C. “Proton inventory of the genomic HDV ribozyme in Mg2+-containing solutions.” J. Am. Chem. Soc., 123, 11333-11334 (2001). [Pubmed]
24. Nakano, S., Proctor, D. J., and Bevilacqua, P. C. “Mechanistic characterization of the HDV genomic ribozyme: Assessing the catalytic and structural contributions of divalent metal ions within a multi-channel reaction mechanism.” Biochemistry, 40, 12022-12038 (2001). [Pubmed]
23. Bevilacqua, P. C. “Making biochemistry required reading.” Chemical & Engineering News. 79, 219 (2001). [pdf]
22. Zheng, X. and Bevilacqua, P. C. “Efficient construction of long DNA duplexes with internal non-Watson-Crick motifs and modifications.” Nucleic Acids Res. 29, E6 (2001). [Pubmed]
2000
21. Zheng, X., and Bevilacqua, P. C. “Straightening of bulged RNA by the double-stranded domain from the protein kinase PKR.” Proc. Natl. Acad. Sci. USA (Track II) 97, 14162-14167 (2000). [Pubmed]
20. Chadalavada, D. M., Knudsen, S. K., Nakano, S., and Bevilacqua, P. C. “A role for upstream RNA structure in facilitating the catalytic fold of the genomic hepatitis delta virus ribozyme.” J. Mol. Biol. 301, 349-368 (2000). [Pubmed]
19. Nakano, S., Chadalavada, D. M., and Bevilacqua, P. C. “General acid-base catalysis in the mechanism of an HDV ribozyme.” Science, 287, 1493-1497 (2000). [Pubmed]
1999
18. Bevilacqua, P. C. “Exploring the possibility of an RNA world.” Book Review of “The RNA World,” 2nd Ed., Gesteland, R. F., Cech, T. R., and Atkins, J. F. Eds. for Nature Struct. Biol., 6, 997 (1999). [pdf]
17. Shu, Z., and Bevilacqua, P. C. “Isolation and characterization of thermodynamically stable and unstable RNA hairpins from a triloop combinatorial library.” Biochemistry, 38, 15369-15379 (1999). [Pubmed]
16. Arnold, J. J., Ghosh, S.K.B., Bevilacqua, P. C. and Cameron, C. “Single-nucleotide resolution of RNA strands in the presence of their RNA complements.” Biotechniques, 27, 450-456 (1999). [Pubmed]
1998
15. Bevilacqua, J. M. and Bevilacqua, P. C. “Thermodynamic analysis of an RNA combinatorial library contained in a short hairpin.” Biochemistry, 37, 15877-15884 (1998). [Pubmed]
14. Szewczak, A. A., Podell, E., Bevilacqua, P. C., and Cech, T. R. “Thermodynamic stability of the P4-P6 domain RNA tertiary structure measure by temperature gradient gel electrophoresis.” Biochemistry 37, 11162-11170 (1998). [Pubmed]
13. Bevilacqua, P. C., George, C., Samuel, C. E., and Cech, T. R. “Binding of the protein kinase PKR to RNAs with secondary structure defects: Role of the tandem A-G mismatch and noncontiguous helixes.” Biochemistry 37, 6303-6316 (1998). [Pubmed]
1996
12. Bevilacqua, P. C. and Cech, T. R. “Minor-groove recognition of double-stranded RNA by the double-stranded RNA-binding domain from the RNA-activated protein kinase PKR.” Biochemistry 35, 9983-9994 (1996). [Pubmed] (pdf)
11. Turner, D. H., Li, Y., Fountain, M., Profenno, L. and Bevilacqua, P. C. “Dynamics of a group I ribozyme detected by spectroscopic methods.” In Nucleic Acids & Molecular Biology, eds Fritz Eckstein & David M. J. Lilley, 10, 19-32 (1996). (pdf)
10. Bevilacqua, P. C., Sugimoto, N. and Turner, D. H. “A mechanistic framework for the second step of splicing catalyzed by the Tetrahymena ribozyme.” Biochemistry 35, 648-658 (1996). [Pubmed] (pdf)
1995
9. Li, Y., Bevilacqua, P. C., Mathews, D. and Turner, D. H. “Thermodynamic and activation parameters for binding of a pyrene labelled substrate by the Tetrahymena ribozyme: Docking is not diffusion controlled and is driven by a favorable entropy change.” Biochemistry 34, 14394-14399 (1995). [Pubmed] (pdf)
1994
8. Bevilacqua, P. C., Li, Y. and Turner, D. H. “Fluorescence-detected stopped flow with a pyrene labeled substrate reveals that guanosine facilitates docking of the 5′ cleavage site into a high free energy binding mode in the Tetrahymena ribozyme.” Biochemistry 33, 11340-11348 (1994). [Pubmed] (pdf)
1993
7. Cech, T. R., Bevilacqua, P. C., Doudna, J. A., McConnell, T. S., Strobel, S. A., Weinstein, L. B. “Mechanism and structure of a catalytic RNA molecule.” In Proceedings of the Robert A. Welch Foundation, 91-110 (1993). (pdf)
6. Turner, D. H. and Bevilacqua, P. C. “Thermodynamic considerations for evolution by RNA.” Invited Chapter in RNA World, Cold Spring Harbor Press, eds Ray Gesteland and John Atkins, 447-464 (1993). (pdf)
5. Bevilacqua, P. C., Johnson, K. A. and Turner, D. H. “Cooperative and anticooperative binding to a ribozyme.” Proc. Natl. Acad. Sci. USA 90, 8357-8361 (1993). [Pubmed] (pdf)
4. Kierzek, R., Li, Y., Turner, D. H. and Bevilacqua, P. C. “5′-Amino pyrene provides a sensitive, non-perturbing fluorescent probe of RNA secondary and tertiary structure formation.” J. Am. Chem. Soc. 115, 4985-4992 (1993). [ACS Link] (pdf)
1992
3. Bevilacqua, P. C., Kierzek, R., Johnson, K. A. and Turner, D. H. “Dynamics of ribozyme binding of substrate revealed by fluorescence detected stopped-flow.” Science 258, 1355-1358 (1992). [Pubmed] (pdf)
1991
2.Bevilacqua, P. C. and Turner, D. H. “Comparison of binding of mixed ribose-deoxyribose analogues of CUCU to a ribozyme and to GGAGAA by equilibrium dialysis: Evidence for ribozyme specific interactions with 2′ OH groups.” Biochemistry 30, 10632-10640 (1991). [Pubmed] (pdf)
1989
1.Sugimoto, N., Tomka, M., Kierzek, R., Bevilacqua, P. C. and Turner, D. H. “Effects of substrate structure on the kinetics of circle opening reactions of the self-splicing intervening sequence from Tetrahymena thermophila: Evidence for substrate and Mg2+ binding interactions.” Nucleic Acids Res. 17, 355-371 (1989). [Pubmed] (pdf)

