Tag Archives: sample clean-up
Desalting intact proteins for the direct infusion ESI MS
Electrospray ionization (ESI) and matrix-assisted laser desorption ionization (MALDI) are both competitive ionization processes in which a more abundant and/or more readily ionizable species will always win. Unless you are studying ionization of Tris, PEG, EDTA or CHAPS in the presence of small amounts of protein, the protein solution submitted for MS analysis should be essentially free of salts, detergents, chaotropic reagents, glycerol and other uninteresting interferences.
“So why not take a 10 mg/mL protein solution in a ‘mild buffer’ and dilute it to the required 0.1 mg/mL; the buffer concentration will be 100 times lower, and it is as good as desalted, right?” Wrong and here’s why:
Dilution is not a solution
Let’s say you have a 10 mg/mL solution of a 10 kDa protein in 10 mM Tris buffer, ‘mild’ enough. This means, you have a 10-fold molar excess of Tris compared to the protein. No matter how much you dilute this solution, the molar ratio of Tris to protein will remain 10:1 while it should be 1:50 or less.
For practical purposes, ‘essentially salt-free’ means that the total molar concentration of interfering species is much lower than that of the protein, e.g. at least 50-fold lower.
Clean-up strategies
There are many ways to clean up a protein solution. In each case, the choice of a protocol will depend on the properties of interfering species and on the size and properties of the protein. For example, detergents can be removed using specialized detergent removal spin columns. PEGs and other charge-neutral contaminants can be washed away by loading the protein onto an ion exchange medium, eluting the protein with a high-salt buffer, and desalting the eluate. Buffer salts, urea, and other small molecules can be removed by either a desalting spin column packed with size-exclusion resin or a centrifugal filter with an appropriate molecular-weight-cut-off (MWCO) membrane. If you need an advice on your protein sample clean-up, I am always happy to help; you know where to find me!
Desalting using centrifugal filters
Centrifugal filters with different MWCO membranes are probably the easiest way to desalt a small volume of protein solution, although there are a few caveats. For example, the regenerated cellulose Ultracel membranes might retain cellulose-binding proteins; in this case, the low protein binding Omega membranes from PALL would be a better choice.
How many spin-dilute cycles are enough to ‘completely’ desalt a protein solution? This is one of those annoying situations where the old trusty triplicate doesn’t always work. Here’s my handy formula to help me decide when to stop desalting and start analyzing:
Example 1.
Let’s use our 10 mg/mL 10 kDa protein in 10 mM Tris buffer and a 3 kDa MWCO Amicon centrifugal filter as an example. The filter has a 0.5 mL capacity, and we can load 50 uL of the protein solution and dilute it with 450 uL of water. After a 30-min centrifugation, the volume is 50 uL, but the concentration of Tris is now 10% (or 0.1) of the initial while the protein concentration is essentially unchanged. So our D, dilution factor for Tris after each dilution+spin is 0.1; our goal, M final is 0.01 mM Tris (100:1 protein:Tris); and our M initial is 10 mM Tris. To find the number of dilute+spin cycles, dust off your scientific calculators:
 After 3 dilute/spin repeats, we will achieve our acceptable concentration of Tris, 100 times lower than the protein concentration.
After 3 dilute/spin repeats, we will achieve our acceptable concentration of Tris, 100 times lower than the protein concentration.
Example 2.
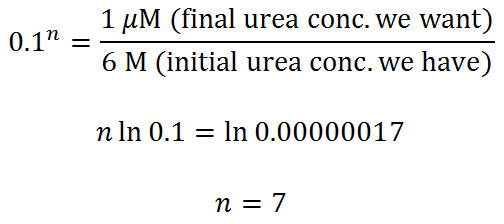
Let’s throw in some urea, dilute the protein to 0.1 mM, and do our calculation again. How many steps (n = ?) do we need to get from 6 M to 1 uM urea (100:1 protein-to-urea ratio) using the same spin column as before?
You probably can get away with 6 dilute/spin repeats, but 7 would be better.
For those of you who hate equations, here’s a picture that illuminates the desalting process using the first example.
“What if my protein is unstable without a buffer?”
In that case, it may not be amenable to the direct ESI MS. However, before throwing up your hands, try to desalt your protein. It might be stable in water at micromolar concentration long enough to be analyzed – I am speaking from experience. You also have an option of using LC ESI MS or submitting your partially desalted protein for MALDI TOF MS.
Ammonium bicarbonate or triethylammonium bicarbonate?
This was a question from one of my blog’s secret readers. Actually, most of the time I feel like I am talking to myself: “Hey Tania, how do you prepare a protein sample for proteolysis?” “Well, Tania, let me show you in a step-by-step tutorial.” No comments, no questions, no pointing out typos, no “thank you, Tania, but there’s a better way to do this”?
Oh well, back to ammonium bicarbonate. This is a volatile salt which breaks down to ammonia, carbon dioxide, and water. Volatile salts are the only salts compatible with MS. Aqueous solutions of ammonium bicarbonate (0.01 – 0.1 M) have pH around 8, the optimal pH for trypsin activity. Ammonium bicarbonate competes with basic amino acids for Coomassie dye, which makes it a great de-staining reagent for the in-gel digestion procedure. All this goodness comes at a very reasonable price – what not to like? Another ammonium salt, triethylammonium bicarbonate (TEAB), is more volatile than ammonium bicarbonate; it is also more expensive. TEAB is a buffer of choice for LC-MS applications: TMT (iTRAQ) amine-reactive labeling, ion-exchange chromatography, protein solubilization (when neutral and acidic pH is undesirable), in-gel digestion, etc.
Detergent removal 2: Affinity resin
To continue our detergent theme, here’s an affinity-based detergent removal method that is faster than the gel-assisted proteolysis but removes only the detergents, leaving the salts and chaotropic agents for you to deal with later. Unlike the gel-assisted method, it works for both proteins and peptides. The gel-assisted method is best suited for proteins, because the small, more soluble peptides are likely to elute out of the gel matrix during the washing.
I use Pierce detergent removal spin columns (0.125 mL format) in my lab, but there are other options available such as Bio-Beads or HyperD, each with its own pros and cons in the business of detergent removal. Pierce also sells a so-called HiPPR detergent removal resin (high protein and peptide recovery) for low-protein-concentration samples. The initial % detergent in such samples must also be low (ca. 1%).
The Pierce resin removes common ionic, nonionic, and zwitterionic detergents from protein and peptide solutions. This oligosaccharide-based affinity resin has a small hydrophobic cavity which creates a microenvironment for a detergent’s nonpolar moiety to enter and form an inclusion complex.
The workflow is simple: (1) centrifuge the column to remove storage buffer, (2) wash 3 times with your favorite buffer (pH 5-10), discarding the buffer each time, (3) add protein or peptide solution and let the resin do its magic for 2-5 min at room temperature, (4) centrifuge to collect your >95% detergent-free sample, i.e. don’t discard the flow-through this time!
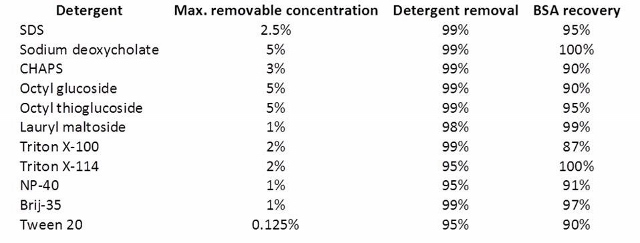
Antharavally and co-workers from Thermo Scientific Pierce Protein Research published a study examining the detergent removal efficiency and protein recovery using their resin under several conditions (doi:10.1016/j.ab.2011.05.013). A table from this reference gives you some idea of the detergent concentrations removable with the Pierce resin.
Samples (0.1 ml containing 0.100 mg BSA + detergent at maximum concentration) were processed through 0.5 ml of Pierce detergent removal resin, and the residual detergent was measured as described in Materials and Methods. Protein concentration was determined by BCA protein assay (Pierce).
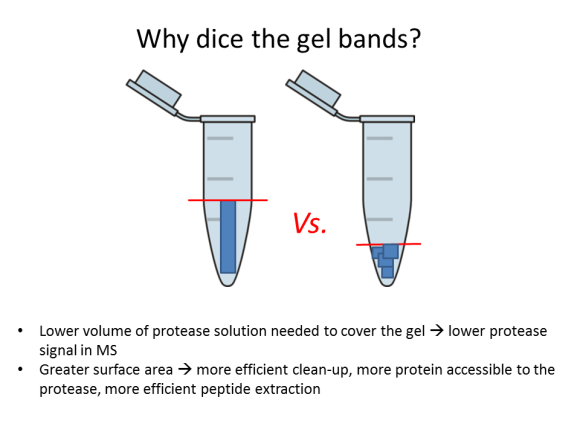
Detergent removal 1: Gel-assisted proteolysis
As promised, here’s a straightforward way to remove detergents, urea, and other LC-MS incompatible nasties from small-volume samples. The literature calls it ‘gel-assisted’ proteolysis. The idea is to entrap the protein solution in a polyacrylamide gel matrix, wash out detergents, salts, and chaotropic agents, and perform in-gel digestion. This technique works great for membrane proteins which are notoriously difficult to dissolve, and it is quite useful for any protein sample clean-up.
For my little demo, I used a 1 mg/mL BSA solution in 2% SDS. The disulfides were reduced with TCEP and alkylated with IAA, after which the protein solution was very thoroughly mixed with a 30% T acrylamide monomer solution. I quickly added 10% APS and TEMED and immediately vortexed and centrifuged this mixture so that the liquid is collected at the bottom of the tube. The polymerization time is very short, a minute or two! I left it to completely polymerize for another 20 min.
Using scalpel I removed the gel plug from the tube and diced it into small pieces. After 6 washes with 8 mg/mL ammonium bicarbonate in 50% acetonitrile, I dried the gel pieces in neat acetonitrile, removed the acetonitrile and added trypsin (see the in-gel digestion tutorial for details).
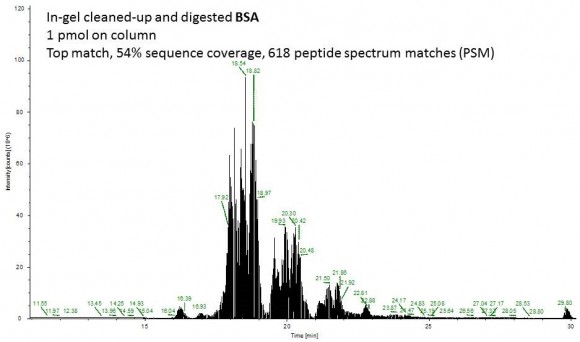
Fast forward to the MS analysis: Since the original BSA solution was very concentrated, I dissolved the peptides in 540 uL of mobile phase and injected 1 uL of this solution (55 ng total protein on column or approximately 1 pmol).
Second example shows an ion chromatogram from a 10-uL IP eluent containing 2% detergent which I cleaned up and digested using this technique.
As always, let me know if you have questions!
References:
doi: 10.1074/mcp.M500138-MCP200
doi: 10.1074/mcp.M800068-MCP200